SpinChainNNN
full name: tenpy.models.spins_nnn.SpinChainNNN
parent module:
tenpy.models.spins_nnntype: class
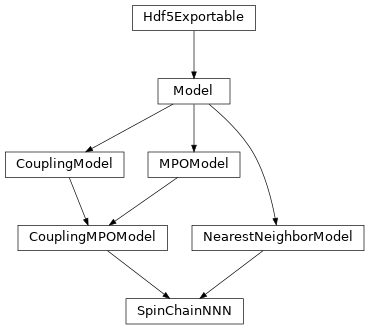
Inheritance Diagram
Methods
|
|
|
Add two-site coupling terms to the Hamiltonian, summing over lattice sites. |
|
Add a two-site coupling term on given MPS sites. |
Add an exponentially decaying long-range coupling. |
|
|
Add a single term to self. |
|
Add multi-site coupling terms to the Hamiltonian, summing over lattice sites. |
|
Add a general M-site coupling term on given MPS sites. |
|
Add onsite terms to |
|
Add an onsite term on a given MPS site. |
Sum of all |
|
Sum of all |
|
Calculate bond energies <psi|H_bond|psi>. |
|
|
Calculate MPO representation of the Hamiltonian. |
|
Calculate the MPO Hamiltonian from the bond Hamiltonian. |
|
calculate H_bond from |
|
Calculate the bond Hamiltonian from the MPO Hamiltonian. |
Shallow copy of self. |
|
Add an external flux to the coupling strength. |
|
|
Repeat the unit cell for infinite MPS boundary conditions; in place. |
Returns the expected saving factor for RAM based on charge conservation. |
|
|
Return a (shallow) copy with extracted segment of MPS. |
|
Initialize a NearestNeighborModel from a model class defining an MPO. |
|
Load instance from a HDF5 file. |
Get list of model-dependent extra default measurements. |
|
|
Modify self in place to group sites. |
Initialize H_MPO (and H_bond) from the terms of the CouplingModel. |
|
|
Initialize a lattice for the given model parameters. |
|
Define the local Hilbert space and operators; needs to be implemented in subclasses. |
|
Add the onsite and coupling terms to the model; subclasses should implement this. |
|
Export self into a HDF5 file. |
Sanity check, raises ValueErrors, if something is wrong. |
|
Return a NearestNeighborModel with same lattice, but trivial (H=0) bonds. |
|
|
Reconstruct Hamiltonian for time-dependent models, potentially (!) in-place. |
Class Attributes and Properties
If True, |
|
class attribute. |
|
Reproducible numpy pseudo random number generator. |
- class tenpy.models.spins_nnn.SpinChainNNN(model_params)[source]
Bases:
CouplingMPOModel,NearestNeighborModelSpin-S sites coupled by (next-)nearest neighbor interactions on a GroupedSite.
The Hamiltonian reads:
\[\begin{split}H = \sum_{\langle i,j \rangle, i < j} \mathtt{Jx} S^x_i S^x_j + \mathtt{Jy} S^y_i S^y_j + \mathtt{Jz} S^z_i S^z_j \\ + \sum_{\langle \langle i,j \rangle \rangle, i< j} \mathtt{Jxp} S^x_i S^x_j + \mathtt{Jyp} S^y_i S^y_j + \mathtt{Jzp} S^z_i S^z_j \\ - \sum_i \mathtt{hx} S^x_i + \mathtt{hy} S^y_i + \mathtt{hz} S^z_i\end{split}\]Here, \(\langle i,j \rangle, i< j\) denotes nearest neighbors and \(\langle \langle i,j \rangle \rangle, i < j\) denotes next nearest neighbors. All parameters are collected in a single dictionary model_params, which is turned into a
Configobject.- Parameters:
model_params (
Config) – Parameters for the model. SeeSpinChainNNNbelow.
Options
- config SpinChainNNN
option summary MPS boundary conditions. Coupling boundary conditions are chosen appropriately.
bc_x (from CouplingMPOModel) in ClockChain.init_lattice
Can be used to force "periodic" boundaries for the lattice, [...]
bc_y (from CouplingMPOModel) in ClockChain.init_lattice
The boundary conditions in y-direction. [...]
What should be conserved. See :class:`~tenpy.networks.Site.SpinSite`.
explicit_plus_hc (from CouplingMPOModel) in CouplingMPOModel
Whether the Hermitian conjugate of the MPO is computed at runtime, [...]
helical (from CouplingMPOModel) in ClockChain.init_lattice
If not ``None``, wrap the lattice into a [...]
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
irregular_remove (from CouplingMPOModel) in ClockChain.init_lattice
If not ``None``, wrap the lattice into a [...]
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
Coupling as defined for the Hamiltonian above.
Length of the chain in terms of :class:`~tenpy.networks.site.GroupedSite`, [...]
lattice (from CouplingMPOModel) in ClockChain.init_lattice
The name of a lattice pre-defined in TeNPy to be initialized. [...]
Lx (from CouplingMPOModel) in ClockChain.init_lattice
The length in x- and y-direction; only read out for 2D lattices. [...]
Ly (from CouplingMPOModel) in ClockChain.init_lattice
The length in x- and y-direction; only read out for 2D lattices. [...]
order (from CouplingMPOModel) in ClockChain.init_lattice
The order of sites within the lattice for non-trivial lattices, [...]
The 2S+1 local states range from m = -S, -S+1, ... +S.
sort_mpo_legs (from CouplingMPOModel) in CouplingMPOModel
Whether the virtual legs of the MPO should be sorted by charges, [...]
- option L: int
Length of the chain in terms of
GroupedSite, i.e. we have2*Lspin sites.
- option S: {0.5, 1, 1.5, 2, ...}
The 2S+1 local states range from m = -S, -S+1, … +S.
- option conserve: 'best' | 'Sz' | 'parity' | None
What should be conserved. See
SpinSite.
- option Jx: float | array
- option Jy: float | array
- option Jz: float | array
- option Jxp: float | array
- option Jyp: float | array
- option Jzp: float | array
- option hx: float | array
- option hy: float | array
- option hz: float | array
Coupling as defined for the Hamiltonian above.
- option bc_MPS: {'finite' | 'infinite'}
MPS boundary conditions. Coupling boundary conditions are chosen appropriately.
- force_default_lattice = True
If True,
init_lattice()asserts that the initialized lattice is (a subclass of) default_lattice- Type:
- init_sites(model_params)[source]
Define the local Hilbert space and operators; needs to be implemented in subclasses.
This function gets called by
init_lattice()to get theSitefor the lattice unit cell.Note
Initializing the sites requires to define the conserved quantum numbers. All pre-defined sites accept
conserve=Noneto disable using quantum numbers. Many models in TeNPy read out the conserve model parameter, which can be set to"best"to indicate the optimal parameters.If you need to initialize more than one site, the function
tenpy.networks.site.set_common_charges()should be helpful.- Parameters:
model_params (dict) – The model parameters given to
__init__.- Returns:
sites ((tuple of)
Site) – The local sites of the lattice, defining the local basis states and operators.optional_species_names (not set | list of str | None) – You should usually just return the (tuple of) sites. However, you can additionally return a list species_names to indicate that the
MultiSpeciesLatticeshould be used.
- init_terms(model_params)[source]
Add the onsite and coupling terms to the model; subclasses should implement this.
- add_coupling(strength, u1, op1, u2, op2, dx, op_string=None, category=None, plus_hc=False)[source]
Add two-site coupling terms to the Hamiltonian, summing over lattice sites.
Represents couplings of the form \(\sum_{x_0, ..., x_{dim-1}} strength[shift(\vec{x})] * OP0 * OP1\), where
OP0 := lat.unit_cell[u0].get_op(op0)acts on the site(x_0, ..., x_{dim-1}, u1), andOP1 := lat.unit_cell[u1].get_op(op1)acts on the site(x_0+dx[0], ..., x_{dim-1}+dx[dim-1], u1). Possible combinationsx_0, ..., x_{dim-1}are determined from the boundary conditions inpossible_couplings().The coupling strength may vary spatially if the given strength is a numpy array. The correct shape of this array is the coupling_shape returned by
tenpy.models.lattice.coupling_shape()and depends on the boundary conditions. Theshift(...)depends on dx, and is chosen such that the first entrystrength[0, 0, ...]of strength is the prefactor for the first possible coupling fitting into the lattice if you imagine open boundary conditions.The necessary terms are just added to
coupling_terms; this function does not rebuild the MPO.- Parameters:
strength (scalar | array) – Prefactor of the coupling. May vary spatially (see above). If an array of smaller size is provided, it gets tiled to the required shape. A single scalar number can be given to indicate a coupling which is uniform across the lattice.
u1 (int) – Picks the site
lat.unit_cell[u1]for OP1.op1 (str) – Valid operator name of an onsite operator in
lat.unit_cell[u1]for OP1.u2 (int) – Picks the site
lat.unit_cell[u2]for OP2.op2 (str) – Valid operator name of an onsite operator in
lat.unit_cell[u2]for OP2.dx (iterable of int) – Translation vector (of the unit cell) between OP1 and OP2. For a 1D lattice, a single int is also fine.
op_string (str | None) – Name of an operator to be used between the OP1 and OP2 sites. Typical use case is the phase for a Jordan-Wigner transformation. The operator should be defined on all sites in the unit cell. If
None, auto-determine whether a Jordan-Wigner string is needed, usingop_needs_JW().category (str) – Descriptive name used as key for
coupling_terms. Defaults to a string of the form"{op1}_i {op2}_j".plus_hc (bool) – If True, the hermitian conjugate of the terms is added automatically.
Examples
When initializing a model, you can add a term \(J \sum_{<i,j>} S^z_i S^z_j\) on all nearest-neighbor bonds of the lattice like this:
>>> J = 1. # the strength >>> for u1, u2, dx in self.lat.pairs['nearest_neighbors']: ... self.add_coupling(J, u1, 'Sz', u2, 'Sz', dx)
The strength can be an array, which gets tiled to the correct shape. For example, in a 1D
Chainwith an even number of sites and periodic (or infinite) boundary conditions, you can add alternating strong and weak couplings with a line like:>>> self.add_coupling([1.5, 1.], u1, 'Sz', u2, 'Sz', dx)
Make sure to use the plus_hc argument if necessary, e.g. for hoppings:
>>> t = 1. # hopping strength >>> for u1, u2, dx in self.lat.pairs['nearest_neighbors']: ... self.add_coupling(t, u1, 'Cd', u2, 'C', dx, plus_hc=True)
Alternatively, you can add the hermitian conjugate terms explicitly. The correct way is to complex conjugate the strength, take the hermitian conjugate of the operators and swap the order (including a swap u1 <-> u2), and use the opposite direction
-dx, i.e. the h.c. ofadd_coupling(t, u1, 'A', u2, 'B', dx)isadd_coupling(np.conj(t), u2, hc('B'), u1, hc('A'), -dx), where hc takes the hermitian conjugate of the operator names, seeget_hc_op_name(). For spin-less fermions (FermionSite), this would be>>> for u1, u2, dx in self.lat.pairs['nearest_neighbors']: ... self.add_coupling(t, u1, 'Cd', u2, 'C', dx) ... self.add_coupling(np.conj(t), u2, 'Cd', u1, 'C', -dx) # h.c.
With spin-full fermions (
SpinHalfFermions), it could be:>>> for u1, u2, dx in self.lat.pairs['nearest_neighbors']: ... self.add_coupling(t, u1, 'Cdu', u2, 'Cd', dx) # Cdagger_up C_down ... self.add_coupling(np.conj(t), u2, 'Cdd', u1, 'Cu', -dx) # h.c. Cdagger_down C_up
Note that the Jordan-Wigner strings for the fermions are added automatically!
See also
add_onsiteAdd terms acting on one site only.
add_multi_coupling_termfor terms on more than two sites.
add_coupling_termAdd a single term without summing over \(\vec{x}\).
- add_coupling_term(strength, i, j, op_i, op_j, op_string='Id', category=None, plus_hc=False)[source]
Add a two-site coupling term on given MPS sites.
Wrapper for
self.coupling_terms[category].add_coupling_term(...).Warning
This function does not handle Jordan-Wigner strings! You might want to use
add_local_term()instead.- Parameters:
strength (float) – The strength of the coupling term.
i (int) – The MPS indices of the two sites on which the operator acts. We require
0 <= i < N_sitesandi < j, i.e., op_i acts “left” of op_j. If j >= N_sites, it indicates couplings between unit cells of an infinite MPS.j (int) – The MPS indices of the two sites on which the operator acts. We require
0 <= i < N_sitesandi < j, i.e., op_i acts “left” of op_j. If j >= N_sites, it indicates couplings between unit cells of an infinite MPS.op1 (str) – Names of the involved operators.
op2 (str) – Names of the involved operators.
op_string (str) – The operator to be inserted between i and j.
category (str) – Descriptive name used as key for
coupling_terms. Defaults to a string of the form"{op1}_i {op2}_j".plus_hc (bool) – If True, the hermitian conjugate of the term is added automatically.
- add_exponentially_decaying_coupling(strength, lambda_, op_i, op_j, subsites=None, op_string=None, plus_hc=False)[source]
Add an exponentially decaying long-range coupling.
\[strength \sum_{i < j} \lambda^{|i-j|} A_{subsites[i]} B_{subsites[j]}\]Where the operator A is given by op_i, and B is given by op_j. Note that the sum over i,j is long-range, for infinite systems going beyond the MPS unit cell. Moreover, note that the distance in the exponent is the distance within subsites.
- Parameters:
strength (float) – Overall prefactor.
lambda (float) – Decay-rate
op_i (string) – Names for the operators.
op_j (string) – Names for the operators.
subsites (None | 1D array) – Selects a subset of sites within the MPS unit cell on which the operators act. Needs to be sorted.
Noneselects all sites.op_string (None | str) – The operator to be inserted between A and B; If
None, this function checks whether a fermionic"JW"string is needed for the given operators; in this case the right op_j acts first.plus_hc (bool) – If True, the hermitian conjugate of the term is added automatically.
Examples
At least for simple enough 1D chains (or ladders), you can use
fit_with_sum_of_exp()to approximate a long-range function with a few sum of exponentials and then add them with this function.>>> def decay(x): ... return np.exp(-0.1*x) / x**2 >>> from tenpy.tools.fit import fit_with_sum_of_exp, sum_of_exp >>> n_exp = 5 >>> fit_range = 50 >>> lam, pref = fit_with_sum_of_exp(decay, n_exp, fit_range) >>> x = np.arange(1, fit_range + 1) >>> print('error in fit: {0:.3e}'.format(np.sum(np.abs(decay(x) - sum_of_exp(lam, pref, x))))) error in fit: 1.073e-04 >>> for pr, la in zip(pref, lam): ... self.add_exponentially_decaying_coupling(pr, la, 'N', 'N')
- add_local_term(strength, term, category=None, plus_hc=False)[source]
Add a single term to self.
The represented term is strength times the product of the operators given in terms. Each operator is specified by the name and the site it acts on; the latter given by a lattice index, see
Lattice.Depending on the length of term, it can add an onsite term or a coupling term to
onsite_termsorcoupling_terms, respectively.- Parameters:
strength (float/complex) – The prefactor of the term.
term (list of (str, array_like)) – List of tuples
(opname, lat_idx)where opname is a string describing the operator acting on the site given by the lattice index lat_idx. Here, lat_idx is for example [x, y, u] for a 2D lattice, with u being the index within the unit cell.category – Descriptive name used as key for
onsite_termsorcoupling_terms.plus_hc (bool) – If True, the hermitian conjugate of the terms is added automatically.
- add_multi_coupling(strength, ops, op_string=None, category=None, plus_hc=False, switchLR='middle_i')[source]
Add multi-site coupling terms to the Hamiltonian, summing over lattice sites.
Represents couplings of the form \(sum_{\vec{x}} strength[shift(\vec{x})] * OP_0 * OP_1 * ... * OP_{M-1}\), involving M operators. Here, \(OP_m\) stands for the operator defined by the m-th tuple
(opname, dx, u)given in the argument ops, which determines the position \(\vec{x} + \vec{dx}\) and unit-cell index u of the site it acts on; the actual operator is given by self.lat.unit_cell[u].get_op(opname).The coupling strength may vary spatially if the given strength is a numpy array. The correct shape of this array is the coupling_shape returned by
tenpy.models.lattice.possible_multi_couplings()and depends on the boundary conditions. Theshift(...)depends on the dx entries of ops and is chosen such that the first entrystrength[0, 0, ...]of strength is the prefactor for the first possible coupling fitting into the lattice if you imagine open boundary conditions.The necessary terms are just added to
coupling_terms; this function does not rebuild the MPO.- Parameters:
strength (scalar | array) – Prefactor of the coupling. May vary spatially, and is tiled to the required shape.
ops (list of
(opname, dx, u)) – Each tuple determines one operator of the coupling, see the description above. opname (str) is the name of the operator, dx (list of length lat.dim) is a translation vector, and u (int) is the index of lat.unit_cell on which the operator acts. The first entry of ops corresponds to \(OP_0\) and acts last in the physical sense.op_string (str | None) –
If a string is given, we use this as the name of an operator to be used inbetween the operators, excluding the sites on which any operators act. This operator should be defined on all sites in the unit cell.
If
None, auto-determine whether a Jordan-Wigner string is needed (usingop_needs_JW()) for each of the segments inbetween the operators and also on the sites of the left operators.switchLR (int |
"middle_i" | "middle_op") – Seeadd_multi_coupling()for details on possible choices.category (str) – Descriptive name used as key for
coupling_terms. Defaults to a string of the form"{op0}_i {other_ops[0]}_j {other_ops[1]}_k ...".plus_hc (bool) – If True, the hermitian conjugate of the terms is added automatically.
Examples
A call to
add_coupling()with argumentsadd_coupling(strength, u1, 'A', u2, 'B', dx)is equivalent to the following:>>> dx_0 = [0] * self.lat.dim # = [0] for a 1D lattice, [0, 0] in 2D >>> self.add_multi_coupling(strength, [('A', dx_0, u1), ('B', dx, u2)])
To explicitly add the hermitian conjugate (instead of simply using plus_hc = True), you need to take the complex conjugate of the strength, reverse the order of the operators and take the hermitian conjugates of the individual operator names (indicated by the
hc(...), seeget_hc_op_name()):>>> self.add_multi_coupling(np.conj(strength), [(hc('B'), dx, u2), (hc('A'), dx_0, u1)])
See also
add_onsiteAdd terms acting on one site only.
add_couplingAdd terms acting on two sites.
add_multi_coupling_termAdd a single term, not summing over the possible \(\vec{x}\).
- add_multi_coupling_term(strength, ijkl, ops_ijkl, op_string, category=None, plus_hc=False, switchLR='middle_i')[source]
Add a general M-site coupling term on given MPS sites.
Wrapper for
self.coupling_terms[category].add_multi_coupling_term(...).Warning
This function does not handle Jordan-Wigner strings! You might want to use
add_local_term()instead.Changed in version 0.10.1: Fix a bug that plus_hc didn’t correctly add the hermitian conjugate terms.
- Parameters:
strength (float) – The strength of the coupling term.
ijkl (list of int) – The MPS indices of the sites on which the operators acts. With i, j, k, … = ijkl, we require that they are ordered ascending,
i < j < k < ...and that0 <= i < N_sites. Indices >= N_sites indicate couplings between different unit cells of an infinite MPS.ops_ijkl (list of str) – Names of the involved operators on sites i, j, k, ….
op_string (list of str) – Names of the operator to be inserted between the operators, e.g., op_string[0] is inserted between i and j.
category (str) – Descriptive name used as key for
coupling_terms. Defaults to a string of the form"{op0}_i {op1}_j {op2}_k ...".plus_hc (bool) – If True, the hermitian conjugate of the term is added automatically.
switchLR (int |
"middle_i" | "middle_op") – Seeadd_multi_coupling()for details on possible choices.
- add_onsite(strength, u, opname, category=None, plus_hc=False)[source]
Add onsite terms to
onsite_terms.Adds \(\sum_{\vec{x}} strength[\vec{x}] * OP`\) to the represented Hamiltonian, where the operator
OP=lat.unit_cell[u].get_op(opname)acts on the site given by a lattice index(x_0, ..., x_{dim-1}, u),The necessary terms are just added to
onsite_terms; doesn’t rebuild the MPO.- Parameters:
strength (scalar | array) – Prefactor of the onsite term. May vary spatially. If an array of smaller size is provided, it gets tiled to the required shape.
u (int) – Picks a
Sitelat.unit_cell[u]out of the unit cell.opname (str) – valid operator name of an onsite operator in
lat.unit_cell[u].category (str) – Descriptive name used as key for
onsite_terms. Defaults to opname.plus_hc (bool) – If True, the hermitian conjugate of the terms is added automatically.
See also
add_couplingAdd a terms acting on two sites.
add_onsite_termAdd a single term without summing over \(vec{x}\).
- add_onsite_term(strength, i, op, category=None, plus_hc=False)[source]
Add an onsite term on a given MPS site.
Wrapper for
self.onsite_terms[category].add_onsite_term(...).- Parameters:
strength (float) – The strength of the term.
i (int) – The MPS index of the site on which the operator acts. We require
0 <= i < L.op (str) – Name of the involved operator.
category (str) – Descriptive name used as key for
onsite_terms. Defaults to op.plus_hc (bool) – If True, the hermitian conjugate of the term is added automatically.
- bond_energies(psi)[source]
Calculate bond energies <psi|H_bond|psi>.
- Parameters:
psi (
MPS) – The MPS for which the bond energies should be calculated.- Returns:
E_bond – List of bond energies: for finite bc,
E_Bond[i]is the energy of bondi, i+1. (i.e. we omit bond 0 between sites L-1 and 0); for infinite bcE_bond[i]is the energy of bondi-1, i.- Return type:
1D ndarray
- calc_H_MPO(tol_zero=1e-15)[source]
Calculate MPO representation of the Hamiltonian.
Uses
onsite_termsandcoupling_termsto build an MPOGraph (and then an MPO).
- calc_H_MPO_from_bond(tol_zero=1e-15)[source]
Calculate the MPO Hamiltonian from the bond Hamiltonian.
- calc_H_bond(tol_zero=1e-15)[source]
calculate H_bond from
coupling_termsandonsite_terms.- Parameters:
tol_zero (float) – prefactors with
abs(strength) < tol_zeroare considered to be zero.- Returns:
H_bond – Bond terms as required by the constructor of
NearestNeighborModel. Legs are['p0', 'p0*', 'p1', 'p1*']- Return type:
list of
Array
:raises ValueError : if the Hamiltonian contains longer-range terms.:
- calc_H_bond_from_MPO(tol_zero=1e-15)[source]
Calculate the bond Hamiltonian from the MPO Hamiltonian.
- Parameters:
tol_zero (float) – Arrays with norm < tol_zero are considered to be zero.
- Returns:
H_bond – Bond terms as required by the constructor of
NearestNeighborModel. Legs are['p0', 'p0*', 'p1', 'p1*']- Return type:
list of
Array
:raises ValueError : if the Hamiltonian contains longer-range terms.:
- coupling_strength_add_ext_flux(strength, dx, phase)[source]
Add an external flux to the coupling strength.
When performing DMRG on a “cylinder” geometry, it might be useful to put an “external flux” through the cylinder. This means that a particle hopping around the cylinder should pick up a phase given by the external flux [resta1998]. This is also called “twisted boundary conditions” in literature. This function adds a complex phase to the strength array on some bonds, such that particles hopping in positive direction around the cylinder pick up exp(+i phase).
Warning
For the sign of phase it is important that you consistently use the creation operator as op1 and the annihilation operator as op2 in
add_coupling().- Parameters:
strength (scalar | array) – The strength to be used in
add_coupling(), when no external flux would be present.dx (iterable of int) – Translation vector (of the unit cell) between op1 and op2 in
add_coupling().phase (iterable of float) – The phase of the external flux for hopping in each direction of the lattice. E.g., if you want flux through the cylinder on which you have an infinite MPS, you should give
phase=[0, phi]such that particles pick up a phase phi when hopping around the cylinder.
- Returns:
strength – The strength array to be used as strength in
add_coupling()with the given dx.- Return type:
complex array
Examples
Let’s say you have an infinite MPS on a cylinder, and want to add nearest-neighbor hopping of fermions with the
FermionSite. The cylinder axis is the x-direction of the lattice, so to put a flux through the cylinder, you want particles hopping around the cylinder to pick up a phase phi given by the external flux.>>> strength = 1. # hopping strength without external flux >>> phi = np.pi/4 # determines the external flux strength >>> for u1, u2, dx in self.lat.pairs['nearest_neighbors']: ... strength_with_flux = self.coupling_strength_add_ext_flux(strength, dx, [0, phi]) ... self.add_coupling(strength_with_flux, u1, 'Cd', u2, 'C', dx) ... self.add_coupling(np.conj(strength_with_flux), u2, 'Cd', u1, 'C', -dx)
- enlarge_mps_unit_cell(factor=2)[source]
Repeat the unit cell for infinite MPS boundary conditions; in place.
This has to be done after finishing initialization and can not be reverted.
- Parameters:
factor (int) – The new number of sites in the MPS unit cell will be increased from N_sites to
factor*N_sites_per_ring. Since MPS unit cells are repeated in the x-direction in our convention, the lattice shape goes from(Lx, Ly, ..., Lu)to(Lx*factor, Ly, ..., Lu).
- estimate_RAM_saving_factor()[source]
Returns the expected saving factor for RAM based on charge conservation.
- Returns:
factor – saving factor, due to conservation
- Return type:
Options
- mem_saving_factor :: None | float
Quantizes the RAM saving, due to conservation laws, to be used by
estimate_simulation_RAM(). By default it is 1/mod, or 1/4 in case of mod=1. However, for some classes this factor might be overwritten, if a better approximation is known. In the best case, the user can adjust this model parameter to enhance the estimate.
- extract_segment(*args, **kwargs)[source]
Return a (shallow) copy with extracted segment of MPS.
- Parameters:
first (int) – See
extract_segment().last (int) – See
extract_segment().enlarge (int) – See
extract_segment().
- Returns:
cp – A shallow copy of self with MPO and lattice extracted for the segment.
- Return type:
Model
- classmethod from_MPOModel(mpo_model)[source]
Initialize a NearestNeighborModel from a model class defining an MPO.
This is especially useful in combination with
MPOModel.group_sites().- Parameters:
mpo_model (
MPOModel) – A model instance implementing the MPO. Does not need to be aNearestNeighborModel, but should only have nearest-neighbor couplings.
Examples
The SpinChainNNN2 has next-nearest-neighbor couplings and thus only implements an MPO:
>>> from tenpy.models.spins_nnn import SpinChainNNN2 >>> nnn_chain = SpinChainNNN2({'L': 20, 'sort_charge': True}) >>> print(isinstance(nnn_chain, NearestNeighborModel)) False >>> print("range before grouping:", nnn_chain.H_MPO.max_range) range before grouping: 2
By grouping each two neighboring sites, we can bring it down to nearest neighbors.
>>> grouped_sites = nnn_chain.group_sites(2) >>> print("range after grouping:", nnn_chain.H_MPO.max_range) range after grouping: 1
Yet, TEBD will not yet work, as the model doesn’t define H_bond. However, we can initialize a NearestNeighborModel from the MPO:
>>> nnn_chain_for_tebd = NearestNeighborModel.from_MPOModel(nnn_chain) >>> isinstance(nnn_chain_for_tebd, NearestNeighborModel) True
- classmethod from_hdf5(hdf5_loader, h5gr, subpath)[source]
Load instance from a HDF5 file.
Same as
from_hdf5(), but handlerng.
- get_extra_default_measurements()[source]
Get list of model-dependent extra default measurements.
Extra measurements for a
Simulation, which depend on the model itself - subclasses should override this method). E.g., aMPOModelshould measure the energy w.r.t. the MPO (Seem_energy_MPO()). However, aNearestNeighborModelshould use the functionm_bond_energies(). The extra measurements are added to the default measurements in_connect_measurements().- Returns:
m_extra_default_list
- Return type:
- group_sites(n=2, grouped_sites=None)[source]
Modify self in place to group sites.
Group each n sites together using the
GroupedSite. This might allow to do TEBD with a Trotter decomposition, or help the convergence of DMRG (in case of too long range interactions).This has to be done after finishing initialization and can not be reverted.
- Parameters:
n (int) – Number of sites to be grouped together.
grouped_sites (None | list of
GroupedSite) – The sites grouped together.
- Returns:
grouped_sites – The sites grouped together.
- Return type:
list of
GroupedSite
- init_H_from_terms()[source]
Initialize H_MPO (and H_bond) from the terms of the CouplingModel.
This function is called automatically during CouplingMPOModel.__init__.
If you use one of the add_* methods of the CouplingModel after initialization, you will need to call init_H_from_terms in the end by yourself, in order to update the H_MPO (and possibly H_bond) representations. (You should get a warning about this… The way to avoid it is to initialize all the terms in init_terms by defining your own model, as outlined in Models.
- init_lattice(model_params)[source]
Initialize a lattice for the given model parameters.
This function reads out the model parameter lattice. This can be a full
Latticeinstance, in which case it is just returned without further action. Alternatively, the lattice parameter can be a string giving the name of one of the predefined lattices, which then gets initialized. Depending on the dimensionality of the lattice, this requires different model parameters.- Parameters:
model_params (dict) – The model parameters given to
__init__.- Returns:
lat – An initialized lattice.
- Return type:
Options
-
option CouplingMPOModel.lattice: str |
Lattice The name of a lattice pre-defined in TeNPy to be initialized. Alternatively, directly a subclass of
Latticeinstead of the name. Alternatively, a (possibly self-defined)Latticeinstance. If an instance is given, none of the further options described here are read out, since they are already given inside the lattice instance!
- option CouplingMPOModel.bc_MPS: str
Boundary conditions for the MPS.
- option CouplingMPOModel.order: str
The order of sites within the lattice for non-trivial lattices, e.g,
'default', 'snake', seeordering(). Only used if lattice is a string.
- option CouplingMPOModel.L: int
The length in x-direction; only read out for 1D lattices. For an infinite system the length of the unit cell.
- option CouplingMPOModel.Lx: int
- option CouplingMPOModel.Ly: int
The length in x- and y-direction; only read out for 2D lattices. For
"infinite"bc_MPS, the system is infinite in x-direction and Lx is the number of “rings” in the infinite MPS unit cell, while Ly gives the circumference around the cylinder or width of th the rung for a ladder (depending on bc_y).
-
option CouplingMPOModel.bc_y:
"cylinder" | "ladder" | "open" | "periodic" The boundary conditions in y-direction. Only read out for 2D lattices. “cylinder” is equivalent to “periodic”, “ladder” is equivalent to “open”.
-
option CouplingMPOModel.bc_x:
"open" | "periodic". Can be used to force “periodic” boundaries for the lattice, i.e., for the couplings in the Hamiltonian, even if the MPS is finite. Defaults to
"open"forbc_MPS="finite"and"periodic"forbc_MPS="infinite. If you are not aware of the consequences, you should probably not use “periodic” boundary conditions. (The MPS is still “open”, so this will introduce long-range couplings between the first and last sites of the MPS!)
-
option CouplingMPOModel.helical:
None| int If not
None, wrap the lattice into aHelicalLatticewith the given int as N_unit_cells.
-
option CouplingMPOModel.irregular_remove:
None| 2D array If not
None, wrap the lattice into aIrregularLatticeremoving the specified sites. To add sites, you need to overwrite the init_lattice method in a custom model.
- logger = <Logger tenpy.models.model.Model (WARNING)>
class attribute.
- Type:
- Type:
An instance of a logger; see Logging and terminal output. NB
- property rng
Reproducible numpy pseudo random number generator.
If you want to add randomness/disorder to your model, it is recommended use this random number generator for reproducibility of the model:
self.rng.random(size=[3, 5])
Especially for models with time-dependence, you can/will otherwise end up generating a new disordered at each time-step!
Options
- random_seed :: None | int
Defaults to 123456789. Seed for numpy pseudo random number generator which can be used as e.g.
self.rng.random(...).
- save_hdf5(hdf5_saver, h5gr, subpath)[source]
Export self into a HDF5 file.
Same as
save_hdf5(), but handlerng.
- trivial_like_NNModel()[source]
Return a NearestNeighborModel with same lattice, but trivial (H=0) bonds.
- update_time_parameter(new_time)[source]
Reconstruct Hamiltonian for time-dependent models, potentially (!) in-place.
For
TimeDependentHAlgorithm, we assume that the model reads out the parameterself.options['time'], and reinitialize/update the model calling this method.- Parameters:
new_time (float) – Time at which the (time-dependent) Hamiltonian should be constructed.
- Returns:
updated_model – Model of the same class as self with Hamiltonian at time new_time. Note that it can just be a reference to self if modified in place, or an entirely new constructed model.
- Return type:
model