Kagome¶
full name: tenpy.models.lattice.Kagome
parent module:
tenpy.models.latticetype: class
-
class
tenpy.models.lattice.Kagome(Lx, Ly, sites, **kwargs)[source]¶ Bases:
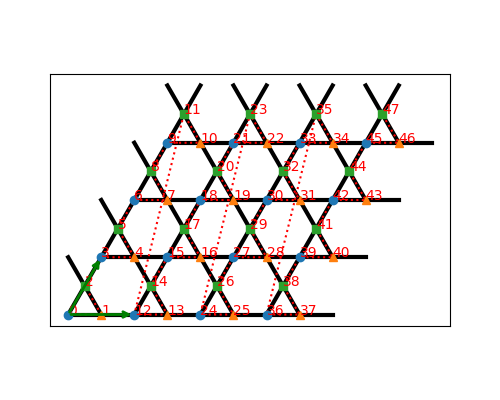
tenpy.models.lattice.LatticeA Kagome lattice.
- Parameters
- Lx, Lyint
The length in each direction.
- sites(list of)
Site The two local lattice sites making the unit_cell of the
Lattice. If only a singleSiteis given, it is used for both sites.- **kwargs :
Additional keyword arguments given to the
Lattice. basis, pos and [[next_]next_]nearest_neighbors are set accordingly.
- Attributes
- nearest_neighbors
- next_nearest_neighbors
- next_next_nearest_neighbors
orderDefines an ordering of the lattice sites, thus mapping the lattice to a 1D chain.
Methods
count_neighbors(self[, u, key])Count e.g.
coupling_shape(self, dx)Calculate correct shape of the strengths for a coupling.
lat2mps_idx(self, lat_idx)Translate lattice indices
(x_0, ..., x_{D-1}, u)to MPS index i.mps2lat_idx(self, i)Translate MPS index i to lattice indices
(x_0, ..., x_{dim-1}, u).mps2lat_values(self, A[, axes, u])Reshape/reorder A to replace an MPS index by lattice indices.
mps_idx_fix_u(self[, u])return an index array of MPS indices for which the site within the unit cell is u.
mps_lat_idx_fix_u(self[, u])Similar as
mps_idx_fix_u(), but return also the corresponding lattice indices.mps_sites(self)Return a list of sites for all MPS indices.
multi_coupling_shape(self, dx)Calculate correct shape of the strengths for a multi_coupling.
number_nearest_neighbors(self[, u])Deprecated.
number_next_nearest_neighbors(self[, u])Deprecated.
ordering(self, order)Provide possible orderings of the N lattice sites.
plot_basis(self, ax, \*\*kwargs)Plot arrows indicating the basis vectors of the lattice.
plot_bc_identified(self, ax[, direction, shift])Mark two sites indified by periodic boundary conditions.
plot_coupling(self, ax[, coupling])Plot lines connecting nearest neighbors of the lattice.
plot_order(self, ax[, order, textkwargs])Plot a line connecting sites in the specified “order” and text labels enumerating them.
plot_sites(self, ax[, markers])Plot the sites of the lattice with markers.
position(self, lat_idx)return ‘space’ position of one or multiple sites.
possible_couplings(self, u1, u2, dx)Find possible MPS indices for two-site couplings.
possible_multi_couplings(self, u0, other_us, dx)Generalization of
possible_couplings()to couplings with more than 2 sites.site(self, i)return
Siteinstance corresponding to an MPS index itest_sanity(self)Sanity check.
-
count_neighbors(self, u=0, key='nearest_neighbors')¶ Count e.g. the number of nearest neighbors for a site in the bulk.
- Parameters
- uint
Specifies the site in the unit cell, for which we should count the number of neighbors (or whatever key specifies).
- keystr
Key of
pairsto select what to count.
- Returns
- numberint
Number of nearest neighbors (or whatever key specified) for the u-th site in the unit cell, somewhere in the bulk of the lattice. Note that it might not be the correct value at the edges of a lattice with open boundary conditions.
-
coupling_shape(self, dx)¶ Calculate correct shape of the strengths for a coupling.
- Parameters
- dxtuple of int
Translation vector in the lattice for a coupling of two operators.
- Returns
- coupling_shapetuple of int
Len
dim. The correct shape for an array specifying the coupling strength. lat_indices has only rows within this shape.- shift_lat_indicesarray
Translation vector from lower left corner of box spanned by dx to the origin.
-
lat2mps_idx(self, lat_idx)¶ Translate lattice indices
(x_0, ..., x_{D-1}, u)to MPS index i.- Parameters
- lat_idxarray_like […, dim+1]
The last dimension corresponds to lattice indices
(x_0, ..., x_{D-1}, u). All lattice indices should be positive and smaller than the corresponding entry inself.shape. Exception: for “infinite” bc_MPS, an x_0 outside indicates shifts accross the boundary.
- Returns
- iarray_like
MPS index/indices corresponding to lat_idx. Has the same shape as lat_idx without the last dimension.
-
mps2lat_idx(self, i)¶ Translate MPS index i to lattice indices
(x_0, ..., x_{dim-1}, u).- Parameters
- iint | array_like of int
MPS index/indices.
- Returns
- lat_idxarray
First dimensions like i, last dimension has len dim`+1 and contains the lattice indices ``(x_0, …, x_{dim-1}, u)` corresponding to i. For i accross the MPS unit cell and “infinite” bc_MPS, we shift x_0 accordingly.
-
mps2lat_values(self, A, axes=0, u=None)¶ Reshape/reorder A to replace an MPS index by lattice indices.
- Parameters
- Andarray
Some values. Must have
A.shape[axes] = self.N_sitesif u isNone, orA.shape[axes] = self.N_cellsif u is an int.- axes(iterable of) int
chooses the axis which should be replaced.
- u
None| int Optionally choose a subset of MPS indices present in the axes of A, namely the indices corresponding to
self.unit_cell[u], as returned bymps_idx_fix_u(). The resulting array will not have the additional dimension(s) of u.
- Returns
- res_Andarray
Reshaped and reordered verions of A. Such that an MPS index j is replaced by
res_A[..., self.order, ...] = A[..., np.arange(self.N_sites), ...]
Examples
Say you measure expection values of an onsite term for an MPS, which gives you an 1D array A, where A[i] is the expectation value of the site given by
self.mps2lat_idx(i). Then this function gives you the expectation values ordered by the lattice:>>> print(lat.shape, A.shape) (10, 3, 2) (60,) >>> A_res = lat.mps2lat_values(A) >>> A_res.shape (10, 3, 2) >>> A_res[lat.mps2lat_idx(5)] == A[5] True
If you have a correlation function
C[i, j], it gets just slightly more complicated:>>> print(lat.shape, C.shape) (10, 3, 2) (60, 60) >>> lat.mps2lat_values(C, axes=[0, 1]).shape (10, 3, 2, 10, 3, 2)
If the unit cell consists of different physical sites, an onsite operator might be defined only on one of the sites in the unit cell. Then you can use
mps_idx_fix_u()to get the indices of sites it is defined on, measure the operator on these sites, and use the argument u of this function.>>> u = 0 >>> idx_subset = lat.mps_idx_fix_u(u) >>> A_u = A[idx_subset] >>> A_u_res = lat.mps2lat_values(A_u, u=u) >>> A_u_res.shape (10, 3) >>> np.all(A_res[:, :, u] == A_u_res[:, :]) True
Todo
make sure this function is used for expectation values…
-
mps_idx_fix_u(self, u=None)¶ return an index array of MPS indices for which the site within the unit cell is u.
If you have multiple sites in your unit-cell, an onsite operator is in general not defined for all sites. This functions returns an index array of the mps indices which belong to sites given by
self.unit_cell[u].- Parameters
- uNone | int
Selects a site of the unit cell.
None(default) means all sites.
- Returns
- mps_idxarray
MPS indices for which
self.site(i) is self.unit_cell[u]. Ordered ascending.
-
mps_lat_idx_fix_u(self, u=None)¶ Similar as
mps_idx_fix_u(), but return also the corresponding lattice indices.- Parameters
- uNone | int
Selects a site of the unit cell.
None(default) means all sites.
- Returns
- mps_idxarray
MPS indices i for which
self.site(i) is self.unit_cell[u].- lat_idx2D array
The row j contains the lattice index (without u) corresponding to
mps_idx[j].
-
mps_sites(self)¶ Return a list of sites for all MPS indices.
Equivalent to
[self.site(i) for i in range(self.N_sites)].This should be used for sites of 1D tensor networks (MPS, MPO,…).
-
multi_coupling_shape(self, dx)¶ Calculate correct shape of the strengths for a multi_coupling.
- Parameters
- dxtuple of int
Translation vector in the lattice for a coupling of two operators.
- Returns
- coupling_shapetuple of int
Len
dim. The correct shape for an array specifying the coupling strength. lat_indices has only rows within this shape.- shift_lat_indicesarray
Translation vector from lower left corner of box spanned by dx to the origin.
-
number_nearest_neighbors(self, u=0)¶ Deprecated.
Use
count_neighbors()instead.
-
number_next_nearest_neighbors(self, u=0)¶ Deprecated.
Use
count_neighbors()instead.
-
property
order¶ Defines an ordering of the lattice sites, thus mapping the lattice to a 1D chain.
This order defines how an MPS/MPO winds through the lattice.
-
ordering(self, order)¶ Provide possible orderings of the N lattice sites.
This function can be overwritten by derived lattices to define additional orderings. The following orders are defined in this method:
order
equivalent priority
equivalent
snake_winding'Cstyle'(0, 1, …, dim-1, dim)
(False, …, False, False)
'default''snake'(0, 1, …, dim-1, dim)
(True, …, True, True)
'snakeCstyle''Fstyle'(dim-1, …, 1, 0, dim)
(False, …, False, False)
'snakeFstyle'(dim-1, …, 1, 0, dim)
(False, …, False, False)
- Parameters
- orderstr |
('standard', snake_winding, priority)|('grouped', groups) Specifies the desired ordering using one of the strings of the above tables. Alternatively, an ordering is specified by a tuple with first entry specifying a function,
'standard'forget_order()and'grouped'forget_order_grouped(), and other arguments in the tuple as specified in the documentation of these functions.
- orderstr |
- Returns
- orderarray, shape (N, D+1), dtype np.intp
the order to be used for
order.
See also
get_ordergenerates the order from equivalent priority and snake_winding.
get_order_groupedvariant of get_order.
plot_ordervisualizes the resulting order.
-
plot_basis(self, ax, **kwargs)¶ Plot arrows indicating the basis vectors of the lattice.
- Parameters
- ax
matplotlib.axes.Axes The axes on which we should plot.
- **kwargs :
Keyword arguments specifying the “arrowprops” of
ax.annotate.
- ax
-
plot_bc_identified(self, ax, direction=-1, shift=None, **kwargs)¶ Mark two sites indified by periodic boundary conditions.
Works only for lattice with a 2-dimensional basis.
- Parameters
- ax
matplotlib.axes.Axes The axes on which we should plot.
- directionint
The direction of the lattice along which we should mark the idenitified sites. If
None, mark it along all directions with periodic boundary conditions.- shiftNone | np.ndarray
The origin starting from where we mark the identified sites. Defaults to the first entry of
unit_cell_positions.- **kwargs :
Keyword arguments for the used
ax.plot.
- ax
-
plot_coupling(self, ax, coupling=None, **kwargs)¶ Plot lines connecting nearest neighbors of the lattice.
- Parameters
- ax
matplotlib.axes.Axes The axes on which we should plot.
- couplinglist of (u1, u2, dx)
By default (
None), useself.pairs['nearest_neighbors']. Specifies the connections to be plotted; iteating over lattice indices (i0, i1, …), we plot a connection from the site(i0, i1, ..., u1)to the site(i0+dx[0], i1+dx[1], ..., u2), taking into account the boundary conditions.- **kwargs :
Further keyword arguments given to
ax.plot().
- ax
-
plot_order(self, ax, order=None, textkwargs={}, **kwargs)¶ Plot a line connecting sites in the specified “order” and text labels enumerating them.
- Parameters
- ax
matplotlib.axes.Axes The axes on which we should plot.
- orderNone | 2D array (self.N_sites, self.dim+1)
The order as returned by
ordering(); by default (None) useorder.- textkwargs: ``None`` | dict
If not
None, we add text labels enumerating the sites in the plot. The dictionary can contain keyword arguments forax.text().- **kwargs :
Further keyword arguments given to
ax.plot().
- ax
-
plot_sites(self, ax, markers=['o', '^', 's', 'p', 'h', 'D'], **kwargs)¶ Plot the sites of the lattice with markers.
- Parameters
- ax
matplotlib.axes.Axes The axes on which we should plot.
- markerslist
List of values for the keywork marker of
ax.plot()to distinguish the different sites in the unit cell, a site u in the unit cell is plotted with a markermarkers[u % len(markers)].- **kwargs :
Further keyword arguments given to
ax.plot().
- ax
-
position(self, lat_idx)¶ return ‘space’ position of one or multiple sites.
- Parameters
- lat_idxndarray,
(... , dim+1) Lattice indices.
- lat_idxndarray,
- Returns
- posndarray,
(..., dim) The position of the lattice sites specified by lat_idx in real-space.
- posndarray,
-
possible_couplings(self, u1, u2, dx)¶ Find possible MPS indices for two-site couplings.
For periodic boundary conditions (
bc[a] == False) the indexx_ais taken moduloLs[a]and runs throughrange(Ls[a]). For open boundary conditions,x_ais limited to0 <= x_a < Ls[a]and0 <= x_a+dx[a] < lat.Ls[a].- Parameters
- u1, u2int
Indices within the unit cell; the u1 and u2 of
add_coupling()- dxarray
Length
dim. The translation in terms of basis vectors for the coupling.
- Returns
- mps1, mps2array
For each possible two-site coupling the MPS indices for the u1 and u2.
- lat_indices2D int array
Rows of lat_indices correspond to rows of mps_ijkl and contain the lattice indices of the “lower left corner” of the box containing the coupling.
- coupling_shapetuple of int
Len
dim. The correct shape for an array specifying the coupling strength. lat_indices has only rows within this shape.
-
possible_multi_couplings(self, u0, other_us, dx)¶ Generalization of
possible_couplings()to couplings with more than 2 sites.Given the arguments of
add_coupling()determine the necessary shape of strength.- Parameters
- u0int
Argument u0 of
add_multi_coupling().- other_uslist of int
The u of the other_ops in
add_multi_coupling().- dxarray, shape (len(other_us), lat.dim+1)
The dx specifying relative operator positions of the other_ops in
add_multi_coupling().
- Returns
- mps_ijkl2D int array
Each row contains MPS indices i,j,k,l,…` for each of the operators positions. The positions are defined by dx (j,k,l,… relative to i) and boundary coundary conditions of self (how much the box for given dx can be shifted around without hitting a boundary - these are the different rows).
- lat_indices2D int array
Rows of lat_indices correspond to rows of mps_ijkl and contain the lattice indices of the “lower left corner” of the box containing the coupling.
- coupling_shapetuple of int
Len
dim. The correct shape for an array specifying the coupling strength. lat_indices has only rows within this shape.
-
test_sanity(self)¶ Sanity check.
Raises ValueErrors, if something is wrong.